caused by a deletion on chromosome 5, in the p15.2 region. The clinical picture involves severe psychomotor and mental retardation, growth retardation, microcephaly and facial dysmorphism.
The G-test evaluates very early (starting from the tenth week) the risk
that the fetus may be affected by a chromosomal disease.
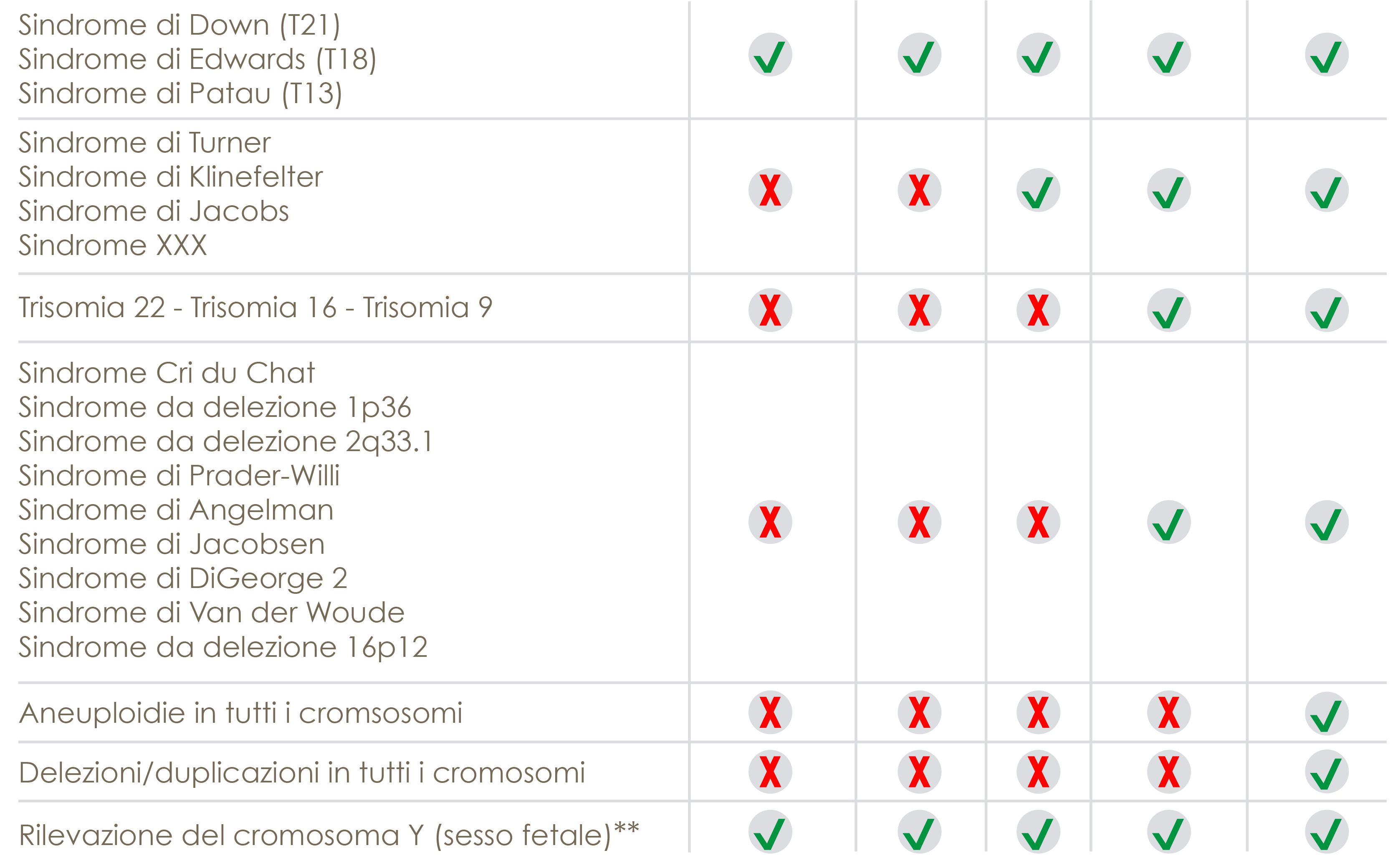
Trisomies
Trisomies are characterized by the presence of a chromosome more than the couple present in the chromosomal kit of a normal individual. The most common at birth is Trisomy 21, associated with Down Syndrome (frequency 1 out of 700 births); Trisomy 18 (Edwards Syndrome, 1 out of 7900 births), Trisomy 13 (Patau Syndrome, 1 out of 9500 births), Trisomy 22, Trisomy 16 and Trisomy 9, usually causing premature abortion, endouterin death, are the rarest. , perinatal or otherwise of short life expectancy.
Abnormalities in the number of sex chromosomes
Aneuploidy of the sex chromosomes are characterized by the absence of a sex chromosome, in the case of Turner's Syndrome (45, X - frequency 1 in 2500 females), or by the presence of an extra sex chromosome, in the case of Klinefelter Syndrome ( XXY, frequency from 1: 500 to 1: 1000 males), of the Jacobs Syndrome (XYY, frequency 1 in 1000 males) and of the XXX Syndrome (frequency 1 in 1000 females).
Deletions
Deletions are unbalanced chromosomal anomalies characterized by the absence of a chromosome trait and, consequently, of the genes located on that fragment. Some deletions cause rare syndromes, which may be associated with: cardiac abnormalities, facial dysmorphism and labiopalatoschisis, eating disorders in early childhood, changes in the functioning of the gastrointestinal tract and immune system, mental retardation or neuro-cognitive impairment . The severity of these clinical manifestations varies, from individual to individual, depending on the size and position of the absent chromosomal fragment.
The G-test evaluates the presence of several deletions responsible for the onset of:
Sindrome da delezione 1p36
caused by a deletion on chromosome 1 in the p36 region. It is associated with moderate to severe intellectual disability, hypotonia, convulsions, pre and post-natal growth retardation, microcephaly, cardiomyopathy, and dysmorphism.
Sindrome da delezione 2p33.1
caused by a deletion on chromosome 2 in the p33.1 region. It is associated with mental retardation and growth, presence of autistic traits, communication difficulties, myocephaly, labio-palatoschisis and facial dysmorphism.
Sindrome da delezione 16p12,2-p11,2
caused by a deletion of chromosome 16. Characterized by developmental delay and cognitive impairment, facial dysmorphism. Orofacial clefts, cardiopathies, short stature, difficulty in feeding and hypotonia can be associated.
Sindrome di Jacobsen da delezione 11q23
associated with growth and psychomotor retardation, facial deformity of the skull, hypertelorism, ptosis, downward palpebral fissures, epicanthus, broad nasal saddle, short nose, V-shaped lips, small ears. Functional abnormalities of platelets, thrombocytopenia or pancytopenia are usually present at birth. Patients often have cardiopathies.
Sindrome di PraderWilli/Angelman 15q11,2
characterized by muscular hypotonia, obesity, difficulty breathing, hypogonadism, intellectual difficulties.
Sindrome Di George 2,12
due to deletion at the level of chromosome 10p14-p13, characterized by heart defects, hypoparathyroidism, T cell immunodeficiency, facial dysmorphism.
Sindrome di Van Der Woude
is a dominant congenital disorder affecting the craniofacial development, in particular with marked labio / cleft palate.
G-Test
Whole Genome Analisys (WGA)
The analysis of the entire genome makes it possible, by applying a proprietary calculation algorithm (FCAPS *), to widen the search for aneuploidies and deletions / duplications to all chromosomes. The WGA Gtest represents the most recent evolution of fetal DNA tests and the overcoming of the obsolete sequencing technologies on single chromosomes (still existing for their low cost). It is therefore possible to perform a non-invasive prenatal screening of the entire chromosomal kit to identify fetal chromosomal abnormalities, even of very small dimensions.
Validation studies of the FCAPS algorithm
● Liu et al., Performance Evaluation of NIPT in Detection of Chromosomal Copy Number Variants Using Low-Coverage Whole-Genome Sequencing of Plasma DNA. PLoS One, 2016
● Chen et al., A method for noninvasive detection of fetal large deletions / duplications. Prenat Diagn, 2013.
* Fetal Copy-number Analysis through Maternal Plasma Sequencing

